 Structural
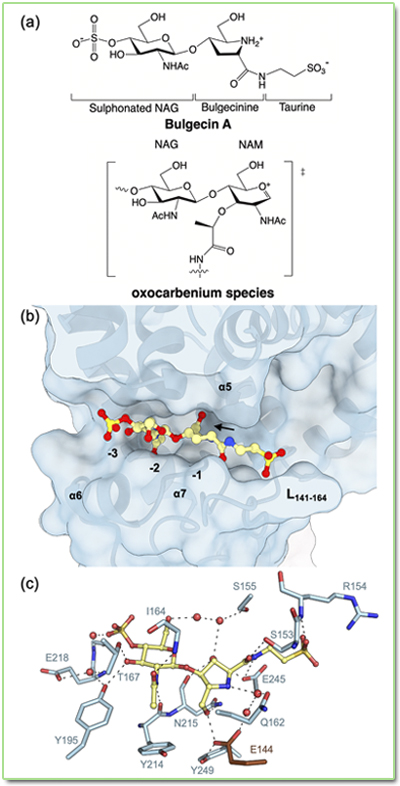
characterization of lytic transglycosylase MltD of Pseudomonas aeruginosa,
a target for the natural product bulgecin A. The natural
inhibitor bulgecin A
potentiates the activity of β-lactam antibiotics through the
inhibition of three lytic transglycosylases in Pseudomonas aeruginosa
(MltD, Slt and MltG). A recent study, led by Juan A. Hermoso from the
Institute of Physical Chemistry “Blas Cabrera” and
published in the International Journal of Biological Macromolecules,
reports three X-ray structures that shed light on the structure of the
multimodular MltD enzyme and provides a detailed molecular recognition
of bulgecin A. These structures reveal a unique combination of a
catalytic module and four cell-wall binding LysM modules: one
unpredicted LysM module tightly attached to the catalytic domain while
others exhibit mobility. A ternary complex structure provides two
independent structures. One delineates the expansive active-site cleft
of MltD by the insertion of a helical region, a hallmark of family 1D
of lytic transglycosylases, providing a handbook explanation of the
endolytic reaction by MltD. The other elucidates the mechanism of the
exolytic reaction, demonstrating how the substrate’s terminal
anhydro-NAM moiety is sequestered at subsite +2. These findings offer a comprehensive
understanding of MltD’s role in cell-wall turnover events,
elucidating both endolytic and exolytic activities, and provide insight
into the molecular recognition of bulgecin A, paving the way for the
development of more effective inhibitors.
International Journal of Biological Macromolecules (2024) (doi: 10.1016/j.ijbiomac.2024.131420)  Unique
cross-link types in Alpha and Betaproteobacteria: A distinctive family
of L,D-transpeptidases catalyzing L-Ala-mDAP crosslinks in Alpha-and
Betaproteobacteria.
The bacterial cell-wall
peptidoglycan is
made of glycan strands cross-linked by short peptide stems. Two
different cross-link types have been long known, 4,3 and 3,3
cross-links, catalyzed by PBP transpeptidases and LD transpeptidases
respectively. A multidimensional work led by Felipe Cava from Umeå
University (Sweden), in collaboration with Juan A.
Hermoso
from the Institute of Physical Chemistry “Blas
Cabrera” has
explored novel cross-link types found in Alpha- and Betaproteobacteria,
adding a new dimension to our understanding of bacterial cell-wall
architecture. The study, published in Nature Communications,
identifies
a LD-transpeptidase from Gluconobacter
oxydans, LDTGo, capable of
generating 1,3 cross-links. LDTGo-like proteins have also been found
among Alpha- and Betaproteobacteria, that lacks LD 3,3 transpeptidases.
A high-resolution structure of LDTGo has been determined, revealing
distinctive features including a Proline-rich region that limits
substrate access, and a cavity for accommodating both glycan and
peptide stem and responsible for the substrate specificity. These
unique properties highlight the diversity of LD transpeptidases.
Furthermore, the study demonstrates the dependence of 1,3 cross-link
formation on substrate availability, involving the function of a DD
endopeptidase. This discovery opens new avenues for understanding
cell-wall integrity and maintenance in bacteria, particularly among
Alpha and Betaproteobacteria.
Nature Communications (2024) 15, 1343 (doi: 10.1038/s41467-024-45620-5)  In
a collaborative effort with
the groups of Lok-To Sham and Luo Min (National Univ. of Singapore) and
Juan A. Hermoso (IQF-CSIC) the machinery of cell division FtsEX:RipC
has been elucidated. The FtsEX complex regulates, directly
or via a protein mediator depending on bacterial genera, peptidoglycan
degradation for cell division. Here we report our investigation of
Mycobacterium tuberculosis FtsEX as a non-canonical regulator with high
basal ATPase activity. The cryo-EM structures of the FtsEX system alone
and in complex with RipC, as well as the ATP-activated state, unveil
detailed information on the signal transduction mechanism, leading to
the activation of RipC. Our findings indicate that RipC is recognized
through a “Match and Fit” mechanism, resulting in
an asymmetric rearrangement of the extracellular domains of FtsX and a
unique inclined binding mode of RipC. This study provides insights into
the molecular mechanisms of FtsEX and RipC regulation in the context of
a critical human pathogen, guiding the design of drugs targeting
peptidoglycan remodeling.
Nature Communications (2023) published online (doi: 10.1038/s41467-023-43770-6) XFELs
to reveal the heterogeneity in M.
tuberculosis β-lactamase inhibition by Sulbactam. This work builds on possibilities
unleashed by mix-and-inject serial crystallography at XFELs. We have
triggered an enzymatic reaction by mixing an inhibitor with enzyme
microcrystals to report, in atomic detail and at room temperature, how
the Mycobacterium
tuberculosis enzyme BlaC is inhibited by sulbactam. Our
results reveal ligand binding heterogeneity, ligand gating,
cooperativity, induced fit, and conformational selection, detailing how
the inhibitor approaches the catalytic clefts and binds to the enzyme
noncovalently before reacting to a trans-enamine.
Nature Communications (2023) (doi: 10.1038/s41467-023-41246-1)  Journal of Biological Chemistry (2023) (doi: 10.1016/j.jbc.2023.105198)  Structure-guided engineering of a receptor-agonist pair for inducible activation of the ABA adaptive response to drought. Abscisic acid (ABA) is a plant hormone that naturally controls the response of plants in drought situations. Based on the atomic structure of ABA receptor proteins, we have designed a synthetic ABA receptor and a small chemical compound that acting together in plants are capable of activating ABA signaling in plants and very efficiently improving their tolerance to drought. Science Advances (2023) 9(10) (doi: 10.1126/sciadv.ade9948) (see video 1) (see video 2) |
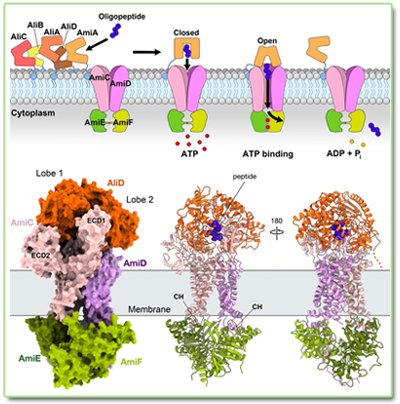 Molecular
and structural basis of oligopeptide recognition by the Ami transporter
system in pneumococci.
ATP-binding cassette (ABC) transport systems are crucial for bacteria
to ensure sufficient uptake of nutrients that are not produced de novo
or improve the energy balance. The cell surface of the pathobiont Streptococcus pneumoniae
(pneumococcus) is decorated with a substantial array of ABC
transporters, critically influencing nasopharyngeal colonization and
invasive infections. Given the auxotrophic nature of pneumococci for
certain amino acids, the Ami ABC transporter system, orchestrating
oligopeptide uptake, becomes indispensable in host compartments lacking
amino acids. The system comprises five exposed Oligopeptide Binding
Proteins (OBPs) and four proteins building the ABC transporter channel.This
work has been led by the groups directed by Juan Hermoso (IQF-CSIC) and
Sven Hammerschmidt (University of Greifswald, Germany), with the
collaboration of the groups led by Isabel Usón (IBMB-CSIC),
Larry McDaniel (University of Mississippi, USA) and
María-José Camarasa (IQM-CSIC). The study,
published now
in PLOS Pathogens
journal,
presents a comprehensive analysis of the initial phase of peptide
transport mediated by OBPs within the pneumococcal Ami permease system.
It discloses a common mechanism for oligopeptide recognition that is
modulated in each OBP to accommodate a diverse array of oligopeptides;
also, a mechanism for membrane transport is proposed. Understanding how
pneumococcus perceives external stimuli and responds to them is
imperative for unraveling the transition from the commensal to the
pathogenic state.
PLoS Pathogens (2024) 20(6): e1011883 (doi:10.1371/journal.ppat.1011883)  IP3K,
the enzyme that metabolizes the second messenger IP3, exhibits unexpected activity on
carbohydrate-based ligands and on those displaying primary hydroxyls in
the reactive position. Inositol 1,4,5-trisphosphate (IP3)
is a second messenger that triggers the release of intracellular Ca2+.
The Ca2+ signals cease when IP3
is metabolized, primarily by the enzyme IP3
3-kinase (IP3K). This enzyme converts IP3 into
Inositol 1,3,4,5-tetrakisphosphate (IP4) and is
crucial for processes such as memory, the immune system, and tumor
progression, making it an attractive target for cancer research.
We have led a study on IP3K in collaboration with Prof. Barry V.L. Potter from the University of Oxford and Dr. Charles A. Brearley from the University of East Anglia. This study broadens our understanding of the biosynthetic capabilities of IP3K beyond its natural substrate, IP3, despite its notable specificity. We have revealed that IP3K exhibits plasticity, conferring tolerance to IP3-derived ligands with modifications, mainly at positions 1 and 3 of the inositol ring. Moreover, the study characterizes the IP3K activity against unexpected ligands, particularly those based on carbohydrates, and those that modify the reactive 3-position from a secondary to primary hydroxyls. To achieve this, we have used X-ray crystallography in combination with multiple techniques including chemical synthesis, fluorescence anisotropy, HPLC or computational docking. These discoveries enhance our understanding of the IP3K family and the inositol polyphosphate metabolism, which are crucial for multiple cell functions. Furthermore, they will aid in the design of selective ligands against different IP3 targets, with potential applications in cancer research. Nature Communications (2024) 15, 1502 (doi: 10.1038/s41467-024-45917-5)  The
genome-organizing protein p6 of Bacillus
subtilis
bacteriophage φ29 plays an essential role in viral development
by activating the initiation of DNA replication and participating in
the early-to-late transcriptional switch. These activities
require the
formation of a nucleoprotein complex in which the DNA adopts a
right-handed superhelix wrapping around a multimeric p6 scaffold,
restraining positive supercoiling and compacting the viral genome. Due
to the absence of homologous structures, prior attempts to unveil p6's
structural architecture failed.
Here, we employed AlphaFold2 to engineer rational p6 constructs
yielding crystals for three-dimensional structure determination. Our
findings reveal a novel fold adopted by p6 that sheds light on its
self-association mechanism and its interaction with DNA. By means of
protein–DNA docking and molecular dynamic simulations, we
have generated a comprehensive structural model for the nucleoprotein
complex that consistently aligns with its established biochemical and
thermodynamic parameters. Besides, through analytical
ultracentrifugation, we have confirmed the hydrodynamic properties of
the nucleocomplex, further validating in solution our proposed model.
Importantly, the disclosed structure not only provides a highly
accurate explanation for previously experimental data accumulated over
decades, but also enhances our holistic understanding of the structural
and functional attributes of protein p6 during φ29 infection.
Nucleic Acids Research (2024) (doi: 10.1093/nar/gkae041)  eLife (2023) 12:e86151 (doi: 10.7554/eLife.86151) New
structural insight into the conformational heterogeneity of NQO1 enzyme
with XFELs. NQO1
is a flavoenzyme essential for the antioxidant defense system,
stabilization of tumor suppressors, and the NAD(P)H-dependent
two-electron reduction of a wide variety of substrates, including the
activation of quinone-based chemotherapeutics. In addition, alterations
in NQO1 function are associated with cancer, Alzheimer's and
Parkinson's disease, which makes this enzyme an attractive target for
drug discovery. The results reported in this article demonstrate the
power of the SFX technique with XFELs to describe the
structure-function relationship in NQO1. We provide important insight
into the conformational heterogeneity of the human NQO1, highlighting
the high plasticity of this enzyme in the catalytic site and hence shed
light on the molecular basis of NQO1 functional cooperativity.
Lab on a Chip (2023) (doi: 10.1039/D3LC00176H)  Nature Communications (2023) 14, Article number: 4095 (doi: 10.1038/s41467-023-39783-w) Mechanistic insights into the regulation of cell wall hydrolysis. The bacterial division is an essential cellular process that involves the formation of a septum made of peptidoglycan. The septum is initially shared between daughters and must be processed to complete division. Septal splitting has long been known to be mediated by enzymes called amidases that are controlled by an activator protein and the ABC-transporter- like complex called FtsEX. However, the mechanism of amidase regulation by this system has remained unclear. In a collaborative effort with the groups of Luo Min and Chris Sam (Univ. Singapore), Thomas Bernhardt (Harvard Univ.) and Juan A. Hermoso (IQF-CSIC), we report the structure of FtsEX in complex with an amidase and amidase activator, revealing how ATP binding to the complex promotes amidase activation and providing structural information that may help target the activation mechanism for the development of cell lysis-inducing antibiotics. Proceedings of the National Academy of Sciences (2023) (doi: 10.1073/pnas.2301897120) See also two short movies: ATP binding leading to PLD restraining and EnvC activation caused by the restraining of PLD upon ATP binding Structural Basis for Cyclosporin Isoform-Specific Inhibition of Cyclophilins from Toxoplasma gondii. Cyclosporin (CsA) has antiparasite activity against the human pathogen Toxoplasma gondii. In a collaborative effort between University of Verona and the IQFR we characterized the functional and structural properties of two cyclophilins from T. gondii, TgCyp23 and TgCyp18.4. While TgCyp23 is a highly active cis−trans-prolyl isomerase (PPIase) and binds CsA with nanomolar affinity, TgCyp18.4 shows low PPIase activity and is significantly less sensitive to CsA inhibition. The crystal structure of the TgCyp23:CsA complex was solved at 1.1 Å resolution showing the molecular details of CsA recognition by the protein, and revealing relevant differences at the CsA-binding site compared to TgCyp18.4. The biochemical and structural data presented herein represents a relevant step toward understanding the molecular mechanisms of the anti-Toxoplasma action of CsA and may be instrumental in the rational design of new therapeutic drugs modulating TgCyp activity ACS Infectious Diseases (2023) (doi: 10.1021/acsinfecdis.2c00566) |










